Summary of key facts
- •
Professor Paul Ehrlich is credited with introducing the concept of cancers as non-self and proposing the concept of tumor immune surveillance in 1909.
- •
Evidence for tumor immunity was suggested by early animal tumor transplantation experiments carried out in nonhomogeneous mouse strains.
- •
The development of congenic inbred murine strains was an essential contribution that enabled the study of human tumor immunogenicity.
- •
TSTAs (tumor transplantation antigens) was the initial term to define tumor antigens. This term was later refined to be TSAs (tumor-specific antigens).
- •
Similarly, tumor-associated transplantation antigens (TATAs) are now called TAAs (tumor-associated antigens).
- •
The human immune system comprises B cells, T cells, and antigen-presenting cells. B cells are the cells responsible for interacting with T cells and antigen-presenting cells to produce highly specific antibodies against antigens.
- •
T cells have subsets; for example, cytotoxic CD8 + T cells (CTLs) and CD4 + T cells.
- •
In 1973, Steinman and Cohen identified a class of antigen-presenting cells termed dendritic cells that function to present antigen in association with MHC class II molecules to CTLs, other T cells, and B cells.
- •
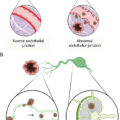
Tumor cells are genetically unstable and a source of altered genes and altered proteins that may become recognized by the host’s immune system as non-self TSAs.
- •
Though SNVs are most common variant occurring in tumor cells, they have not proved to be a very good source of neoantigens. Indels (insertions and deletions) are the second most common type of variant occurring in tumor cells.
- •
Studies indicate that TSAs originating from genomic variants occurring in noncoding regions of the tumor genome (alternative reading frames, antisense coding segments, variants altering splicing in formation of mRNA, 5′ untranslated regions, and long noncoding RNAs) are proving to be a more important source of actionable TSAs.
- •
TAAs actually represent a form of wt peptide but are found in much greater abundance (overexpression) as a result of epigenetic alterations in tumor cell DNA methylation and/or alterations of chromatin structure resulting in shifts in gene expression. TAAs also include cancer germline antigens.
- •
The proteasome and its chamber located in the cytosol function as site of proteolysis to generate short antigenic peptides from the altered proteins suitable for restricted MHC class I presentation. The proteasome can ligate distal peptides together (transpeptidation).
- •
Peptides are imported into endoplasmic reticulum where they are noncovalently bound by the peptide-loading complex (involving multiple proteins) to the MHC binding groove.
- •
The MHC heterodimer is composed of genetically polymorphic α and β chains and the peptide grove of MHC class I is typically closed at both ends, thus accepting only short peptides.
- •
Neoantigen peptide binding is controlled by a series of four to five pockets into which amino acid side chains of the peptide selectively dock.
- •
Bioinformatics tools, increasing computational power, and a growing number of critical data sets supporting machine learning all promise a future enabling antigen identification; prediction for optimal antigen processing, selective MHC binding, successful TCR presentation, and adaptive immune response; and, most important, the suitability as a therapeutic target.
Introduction
One of the great experiences in my career was the time spent at the famous Karolinska Institute, Stockholm, Sweden. I arrived there in 1970 during a break in my surgery training and with very little experience as a student of immunology. One of the special benefits of joining the Möller laboratory on the Karolinska campus was the proximity to the laboratory of the famous immunologist and cancer researcher Professor George Klein, who headed the Karolinska Institute’s Department of Tumor Biology. My mentors, Erna and Göran Möller, had both trained under Professor Klein, so there were lots of opportunities for cross-lab interactions. This was the very early days of tumor immunology and just the beginning of the recognition that the immune system involved two major, functionally unique, immune cells, T cells and B cells. Of course, this was long before the identification of T-cell subtypes such as tumor-directed cytotoxic T lymphocytes (CD8 + CTLs) and the defining of the unique functions of these cells in the adaptive antitumor immune response. The exciting story of immuno-oncology today is very much the history of our evolving understanding of the intricacies of the human immune system and its response to foreign “antigens.”
The primacy concept underpinning the development of our newest weapon against cancer—immunotherapy—is the understanding that when normal cells undergo genetic transformation to the cancer phenotype and acquire their unique capacity for uncontrolled division, the invasion of normal tissues, and the spread to distant sites, they have also acquired non-self-antigens. Tumor cells are, by their very nature, quite genetically unstable and have been shown to express between 50 and 1000 missense mutations. Theoretically, this array of genetic alterations should result in numerous antigenic differences from self and be adequate to trigger a reaction within the host’s immune system to destroy these abnormal cancer cells in their earliest state of development. A process often referred to as immunosurveillance.
The initial introduction of the hypothesis of immune surveillance is most often credited to the German Professor Paul Ehrlich. As early as 1909, Professor Ehrlich reasoned that without the host’s immune system monitoring the development of abnormal cancerous cells, “so konnte man vermuten, dass das Karzinom in einer geradezu ungeheurlichen Frequenz aufteten wurde,” which translates as “one could therefore suspect that the carcinoma would occur with an almost unbelievable frequency.” , However, the state of our knowledge at that time regarding the intricacies of the immune system, as well as the limitations of available scientific tools, made it difficult to validate his hypothesis. Thus, over the years the concept of “host immune surveillance” remained controversial and in fact, does so even today. (See also Chapter 6 , Tumor Immune Surveillance.)
Early experiments involved the simple grafting of fresh pieces of animal tumor tissue into mice or rats in an effort to immunize the animal to a subsequent tumor challenge. Beginning in the early years of the 20th century, scientists repeatedly observed what appeared to be immunologic differences between tumor tissues and normal tissues. Much of this early work involved the use of experimental tumors occurring spontaneously as well as those induced by chemicals such as methylcholanthrene and tumors produced by DNA and RNA oncogenic viruses. Despite somewhat variable findings, there was general recognition that tumor antigens, to be defined as such, needed to be present only in the tumor cells and not present in normal cells. Further, immunity to this tumor antigen brings with it the ability to destroy the tumor cell.
Lumsden is most often given credit for being the first to use an early attempt at developing an inbred mouse strain in his experiments to study the immune response to tumor tissue. He used an “inbred” strain of mice obtained from Lashoploeb’s laboratory in Buffalo that were more than 32 generations. For his experiments, Lumsden used a mouse tumor that had developed spontaneously. Unfortunately, of 170 mice tested, only 3 proved resistant to the tumor.
In 1935, Besredka and Gross reported a series of mouse experiments in which a small amount of a tumor cell suspension was injected intradermally. This resulted in tumor growth which soon regressed and appeared to make the mice resistant to subsequent inoculation of the tumor. Gross later reported a similar set of experiments but using the inbred mouse strain C3H developed by over 20 years of brother–sister mating. Gross used a methylcholanthrene tumor that had been induced originally in C3H mice. In these experiments, the immunized group was resistant to tumor challenge but, interestingly, Gross noted that this resistance to tumor challenge could be overcome by using a larger challenge dose of tumor cells. ,
Sjögren is most often credited with proposing that the antigens unique to tumor cells be designated as tumor-specific transplantation antigens (TSTAs). , This was based on the observation that tumors in mice induced by a small DNA virus contained a common immune response-inducing antigen. TSTAs were capable of being identified on the surface of tumor cells by antibodies they induced in the host. Though they were capable of killing tumor cells in a complement-dependent fashion in vitro, it soon became apparent that such antibodies directed at the surface TSTAs of carcinomas and sarcomas did not protect the host animal from tumor challenge.
The majority of these reported efforts to demonstrate tumor immunogenicity or the existence of TSTAs, dating from the turn of the century and through the 1940s and early 1950s, were carried out in genetically nonhomogeneous mouse strains. Therefore in almost all instances they were most often the result of normal tissue transplantation antigens also present on tumor cells rather than any unique tumor antigen. These observations by the early “giants of immunology”—R. T. Prehn, G. Klein, H. O. Sjögren, K. E. Hellström, and others—demonstrated to varying degrees evidence that cancers possessed non-self unique antigens. Much of this early work to demonstrate TSTAs, however, was rewritten once immunologists, perhaps influenced by Drosophila geneticists of the time, recognized the necessity of developing truly inbred strains of mice to study mammalian genetics and immunology.
In the early 1970s after returning from the Karolinska Institute, I was also privileged to be mentored by Donald C. Shreffler, an immunogeneticist and professor in the Department of Human Genetics at the University of Michigan. Shreffler was a recognized pioneer in researching the genes, including their functional roles, that comprised the murine major histocompatibility and the human leukocyte antigen (HLA) system. Don and his lab were especially recognized for developing numerous, very precious inbred congenic and recombinant H2 complex murine strains as well as strain-specific antisera to the proteins of the H2 locus. It was during these years that the time invested in developing truly isogenic mouse strains began to bear fruit and define the critically dominant genes governing tissue transplantation termed the major histocompatibility complex (MHC) and the processes of antigen recognition. Understanding the role of MHC in antigen presentation was a crucial step in beginning to define and characterize true tumor-specific antigens (TSAs).
During this time, immunology evolved to the point at which it was evident that there existed two principal immunologically competent cell types, the antibody-producing B lymphocytes and the thymus-derived lymphocytes termed T cells. , As a result, experiments demonstrated that T cells could be sensitized to TSTA and alone, in either in vitro or in vivo experimental models, could be shown to destroy targeted tumor cells by a process known as cell-mediated killing . Thus, there was beginning to be ample evidence to support the presence of TSTAs in animal models, but it remained to demonstrate their presence in human tumors. Because tumor transplantation experiments were limited to animals, the analogous antigenic proteins in humans were more appropriately termed tumor-specific antigens . TSAs are defined as being specific to the tumor and, as a result, are not proteins or oligopeptides normally present in the nontumor tissues of the body.
TSAs can result from somatically mutated genes and structural genomic variants occurring during tumor evolution and, as such, produce structurally novel tumor proteins. They can also arise secondary to exposure of the cell’s DNA to viral genomes and to carcinogens. A second class of tumor antigens have been classified as being tumor-associated transplantation antigens (TATAs). TATAs, later termed simply tumor-associated antigens (TAAs), are normal cellular proteins that, when overexpressed by growing tumors, become antigenic in terms of immune recognition and no longer elicit complete immunologic tolerance in the patient. Several examples of the presence of human TAAs can be found in breast cancer, ovarian cancer, and prostate cancer, as well as proteins normally present in skin melanocytes at low levels. Melanoma has been the most intensely studied human tumor in terms of TSAs and TAAs, perhaps because it is an example of a human malignancy that can undergo spontaneous remission (see Tumor-associated antigens ).
Identification of human tumor antigens
I have often thought that it was the surgeons’ feverish efforts to successfully replace diseased human organs with transplanted heterologous tissues that, in many ways, drove a new generation of young scientists to enter the field of immunology. Renal transplantation presented the human model for developing the needed methods of immune suppression and thus the need to understand the mechanisms underlying the complex processes of tissue rejection. I still recall my excitement as a young surgeon seeing the newly transplanted kidney produce urine even before completing the implantation of the ureter into the bladder and, as a result, markedly changing the life of the recipient. Organ transplantation and the challenges of immunosuppression really opened up the field of immunology and attracted many young scientists to study immunology.
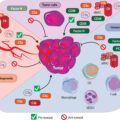
As a result, advances in the field of immunology, driven by organ transplantation, had a tremendous effect on the study of the immune system’s interaction with a progressing human cancer. Today, the central efforts in this new era of cancer immunotherapy are directed at generating an enhanced adaptive immune response, the end result of which is a focused destruction of the cancer cells by cytolytic T lymphocytes (CTLs), while avoiding significant unwanted immune-based toxicities. This adaptive response, whether CTL-based, antibody-directed immune checkpoint inhibition, vaccines, oncolytic viruses, or some combination of these, requires the recognition of unique TSAs/TAAs and a process for their presentation that stimulates an adaptive immune response against the cancer.
A brief reflection on important historic events regarding TSAs and TAAs
It is important perhaps to pause a moment and reflect on the tools of the field and our historical understanding of the immune response to a foreign antigen, whether of an infectious origin, an evolving tumor, or a transplanted organ. As noted earlier, the demonstration of the presence of the T lymphocyte as a critical cellular component of the immune system was a landmark step forward in the mid-1960s. I remember very clearly the ability to use “antitheta” antibodies to separate mouse T cells from B lymphocytes for use in our in vitro antigen presentation experiments, as well as the challenge purifying a rabbit anti-B cell antibody during my days in the Möller lab.
It soon became clear that the TSAs were recognized by the T cells often to be found in the tumor itself and in the tumor-draining lymph nodes. The recognition that it was T cells interacting with antigenic determinants on the tumor cells and the resultant tumor cell destruction marked the beginning of characterizing the subclasses of T lymphocytes, including CD8 + cytolytic T cells (CTLs) and eventually CD4 + T cells.
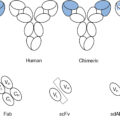
In 1973, Ralph Steinman and Zanvil Cohn reported on the identification of a novel “glass and plastic” adherent cell found in the skin and peripheral lymphoid tissues termed dendritic cells (DCs). These cells soon became recognized as a critical cellular component of the immune system response to foreign antigen. Today, a number of morphologically similar cells have been defined as DCs with a diversity of functions. The classic DC has the ability to be an immunostimulatory cell in response to specific stimuli and in this higher state of differentiation to express high levels of peptide-bound MHC class II molecules that can interact directly with T cells. DCs are capable of recognizing foreign antigen materials—bacteria, viruses, toxins, and TSAs (tumor-derived peptides and lipids)—and through their own proteosome converting them to small peptide fragments to be presented in association with MHC class II molecules for adaptive immune response presentation. The classic DC, we now know, is also capable of interacting with and transmitting information to B cells, as well as other immune system cells such as CD4 + T cells and natural killer (NK) cells (for an in-depth review, see Cabeza-Cabrerizo et al. ). Another important advance was the development by Kohler and Milstein in 1975 of the ability to generate immortalized B cells and to create hybridoma cell lines that would produce an abundance of specific monoclonal antibodies. These murine monoclonal antibodies were used early on to identify TSAs on mouse tumors and eventually on some human cancers. The evolution of this discovery has today become a central feature of immuno-oncology.
In 1989, Lurquin and colleagues reported a series of experiments demonstrating that a peptide derived from a normal self-protein that had become mutated in cancer cells could be recognized by CTLs. This observation has been expanded experimentally over the years to define the process of antigen presentation and T-cell receptor recognition. Recognition of the antigenic peptide begins in the cytosol of the tumor cell where genetically altered self-proteins occur as part of the tumor cell’s cancerous progression. These altered proteins are cleaved into small peptides that are presented at the surface of the tumor cell, complexed in the binding cleft of the MHC class I molecules where they can be presented and recognized by the T-cell receptors (TCRs).
These MHC class I–associated peptides, often referred to as MAPs, are key to the process of CD8 + T-cell maturation and their ability to functionally discriminate between normal cells (self) and those cells that are infected or cancerous (non-self). Thus, the vast majority of TSAs are products of normal cellular proteins that have been genetically altered during tumor initiation and progression and subsequently processed to be non-self-MAPs. This intracellular process of the presentation of MAPs on the cell surface can occur in virtually all cells of the body and is often referred to as the immunopeptidome processing mechanism. , It should be remembered that MHC class I alleles are highly polymorphic and, therefore, each allotype has a unique MAP peptide-binding motif. It is important to note that in contrast to MAPs, the process of antigen presentation by MHC class II determinants is the special activity of the classical antigen-presenting cells: the macrophages, dendritic cells, and B cells.
Two caveats appear worth noting. First, though much of the search for TSAs has naturally focused on the genetic alterations occurring in the exome of the gene, it is increasingly apparent that a much more common occurrence is MAPs derived from the unmutated peptides of cancer-specific epigenetic and splicing alterations. , The existence of MAPs derived from noncoding regions of the genome greatly increases the number of TSAs potentially recognizable by CD8 + T cells from roughly 2% to perhaps as great as 75%. Secondly, cancer cells have proven to be rather poor antigen-presenting cells and therefore the response to TSAs is significantly dependent on cross-presentation by DCs. If this DC presentation does not occur in sufficient robustness, then many potential TSAs are not immunologically active. ,
As indicated earlier in this chapter, antigenic neoepitopes as in TSAs unique to the tumor (or a class of tumors such as melanoma) must be recognized by T cells through the T-cell surface receptor (TCR). What makes this MHC class I peptide complex a valuable therapeutic tool in cancer immunotherapy is the tremendous antigenic epitope binding diversity of these T-cell receptors—a receptor-binding diversity similar to that of B cell–produced antibodies. The generation of the immunogenic neoepitope has its origin in the cytosol, where in the case of a cancer cell, altered self-proteins are cleaved by the proteasome and aminopeptidases into peptide fragments. The peptides are then translocated in the endoplasmic reticulum (ER) via a transporter associated with antigen processing and further resolved by enzymes (ER aminopeptidases) to reach a size of ∼8 to 10 amino acids. These neoepitope peptides are then loaded into the specific peptide cleft of an MHC class I molecule, and if the complex is stable, it is exported as a MAP complex to the tumor cell surface where it can be recognized by TCRs of the T cell. A more detailed description of the intracellular processes operative in creating the short neoantigen peptides for MHC class I binding is presented later in this chapter in the section Tumor antigens—Intracellular processing and MHC presentation .
The origins of targetable tumor-specific antigens—today’s science
The rapid evolution of affordable high-throughput sequencing—based assays, advances in mass spectroscopy, the National Cancer Institute’s Cancer Genome Atlas (TCGA) project, the accumulating number of critical databases, and advances in computational biology have resulted in exciting advances in the field of tumor biology. These tools of science are providing new insights into tumor-specific antigenicity and the potential for developing novel immunotherapies. This rapid progress since 2005 has led to some confusion in the field regarding the true definition of tumor antigens and their actual potential as useful targets. This section is designed to add some definition and clarity to our understanding of the various sources of tumor antigens arising in the genetically unstable cancer cell.
TSAs can be subdivided broadly into two groups: those derived from exon or nonexon mutated/altered DNA gene sequences (mTSAs) and those derived from aberrant expression of transcripts not expressed in nonmalignant tissues, including in medullary thymic epithelial cells, which provide a source of antigen tolerance. A second group of TSAs are derived from human endogenous retroelements (EREs) that comprise approximately 42% of our genome. Much of this part of our genomic DNA results from the integration of transposable elements into the genome millions of years ago (retroviruses) that, through genetic evolution, have lost their ability to be expressed. Aberrantly expressed EREs, however, can occur in the genomically unstable cancer cell via epigenetic dysregulation of the cancer genome such as H3K27me3 loss. You will see these latter TSAs frequently denoted as aeTSA. ,
Single nucleotide variant neoantigens
Tumor cells as altered self are by their very nature genetically unstable and undergo a series of genetic and epigenetic alterations during tumor growth and metastasis. Single nucleotide variants (SNVs) are the most common genetic alteration even in normal cells and, as such, have long been considered an excellent initiator of novel tumor TSAs. These neoantigens and their resultant MAPs have been considered a source of potential antitumor T-cell targets. Nonsynonymous SNVs are most commonly the result of simple DNA replication errors during abnormal tumor cell division and are sometimes positively selected. SNVs can also result from genetic alterations in the tumor cell that damage critical enzymes essential for faithful DNA damage repair. SNVs can, of course, result from ongoing exposure to external mutagens during cancer treatment.
As therapeutically promising as SNV-generated TSAs may appear, it has proved much more difficult to identify suitable targets for the development of novel immunotherapies. As pointed out by Minati and colleagues at the Université de Montreal, Canada, in their excellent review, except perhaps for highly mutated tumors such as melanoma and lung, “all studies based on whole-exome or ribonucleic acid sequencing (WES and RNAseq respectively) combined or not with mass spectrometry (MS) analysis, could only identify a very limited number of SNV-derived MAPs.” ,
It appears, therefore, that sources of neoantigens other than those originating from exome SNVs will be more critical to the design of future immunotherapies. In fact, in mass spectroscopy studies by Laumont and colleagues, some 90% of the identified TSAs in two mouse cancer cell lines and seven primary human tumors were unexpectedly found to be derived from noncoding regions of the genome. These potentially more important sources for actionable TSAs will be discussed in later sections of this chapter. ,
Insertions and deletions (indels)
Insertions and deletions (indels), small nucleotide (nt) insertions or deletions, represent the second most common type of genomic alteration found in cancer cells but, as will be discussed, are by their nature more likely to prove immunogenetic than the more common SNVs. Single nt insertions or deletions are most common and are estimated to account for ∼68% of indels. When nt insertions or deletions occur as in-frame indels, they are, as expected, less likely to be significantly immunogenic. In contrast, those that result in frameshift variants cause the protein produced to be truncated or to be otherwise significantly altered. This can generate peptide fragments and eventual MAPs that are highly immunogenic. Frameshift indels create a novel open reading frame with the potential of producing a large number of peptide fragments that are highly distinct from self and therefore potentially attractive tumor neoantigens.
Studies by Turajlic and colleagues searched TCGA for whole-exome sequence data of 5777 solid tumors across 19 cancer organ sites. They found, for example, that the pattern of indel mutations across the various subclasses of kidney cancer had the highest pan-cancer burden of indel mutations and were enriched for frameshift-based mutant-specific neoantigens. They also reported that in their studies the indel mutation number in cutaneous melanoma was significantly associated with the presence of an effective response to checkpoint inhibitors. Renal cell cancers and melanoma, interestingly, have historically been recognized as immunologically responsive and to have a significance occurrence of spontaneous regression and are known to have a more abundant presence of tumor-infiltrating lymphocytes (TILs). TILs are often seen as a predictor of tumor immune responsiveness. Frameshift-based mutations have also been reported to be more abundant in tumors harboring impaired functions of homologous recombination and DNA mismatch repair.
TSAs derived from altered mRNA splicing
In discussing the tumor cell’s dysregulation of the normal processes involved in the generation of cellular proteins, it is relatively easy to understand how the instability of such a complex intracellular process as messenger RNA (mRNA) splicing could become an important source of TSAs. In order for a gene to be correctly translated into protein, the mRNA transcribed by RNA polymerase must first be processed and spliced together to remove all noncoding regions (introns). In this multistep nuclear process, the splicing together of the exons requires many highly regulated enzymatic proteins and small RNA molecules. Taken together, the machinery for this process is termed the spliceosome . Thus, the mRNA splicing process involves the formation of a highly dynamic and complex spliceosome and a catalytic process to accomplish the required sequence rearrangements. Any dysregulation of these complex mechanisms within the transformed tumor cell is a potential origin for TSAs.
As the intricacies of transcription and translation have been increasingly defined, anticancer therapeutic efforts have been initiated that are directed at correcting or inhibiting core components of the spliceosome in the cancer cell. The future holds great promise that research into the dysregulation of the mRNA splicing mechanisms in malignancy will provide not only potential new TSAs but opportunities for the development of future anticancer therapeutics (for an in-depth review, see Blijlevens et al. ). In addition to targeting abnormalities of the spliceosome itself, alterations in the splicing process of the tumor cell have the potential to result in novel transcripts not found in the patient’s normal cells. These cancer-specific transcripts can be translated by the tumor cell into new isoforms of the protein, resulting in immunogenic TSAs. Aberrantly spliced mRNAs, whether arising from cis -acting splice junction mutations or transacting spliceosome dysregulation, have the potential to result in cancer-specific neojunctions. , The nature of the cancer-specific messenger neo junction can significantly affect the function and structural formation of the resultant protein and its potential immunogenicity as a suitable TSA. It is believed that altered splicing events—whether cis -acting or trans -acting—are more likely to result in antigenic peptides capable of binding to MHC class I molecules for antigen presentation than those TSAs resulting from SNVs. Beyond a potentially novel TSA as a result of aberrant splicing, the resultant protein isoforms with altered function may also affect the tumor in a variety of ways. For example, the resultant proteins may make the tumor more resistant to therapies—both immunotherapies and chemotherapy.
Aberrant transcription TSAs (pre-mRNA alternative splicing)
We have discussed at some length the generation of actionable TSAs as a result of altered splicing events in the genomically unstable tumor cell. Our discussion focused largely on the trans- acting splicing mutations that target the enzymatic proteins and small RNA factors of the spliceosome as a source for generating altered tumor-specific antigenic peptides with potential for T-cell recognition. The pre-mRNA stage in the nucleus can also be a source for the formation of TSAs. The pre-mRNA stage involves alternative choices of exon assembly secondary to mutated splice sites, which result in multiple (mRNA) isoforms being produced from a single gene.
The process of alternative splicing in a normal cell is the major way in which a single gene can create protein diversity. Alternative splicing, in general, occurs as a result of one of five basic actions: (1) exon skipping, (2) use of different options at the 5′ splice site, (3) alternatives at the 3′ splice site, (4) as the result of mutually exclusive exons; and (5) as the result of intron sequence retention. In addition, the family of a specific gene’s mRNA isoforms can be further expanded by a process termed RNA editing . RNA editing is the mechanism by which an RNA transcript can undergo site-specific nt change. An example of the most common mRNA editing is when adenosine deaminase catalyzes the conversion of adenosine to inosine (A to I). A to I RNA editing is associated with cancer progression and impaired patient survival ( Fig. 5.1 ).
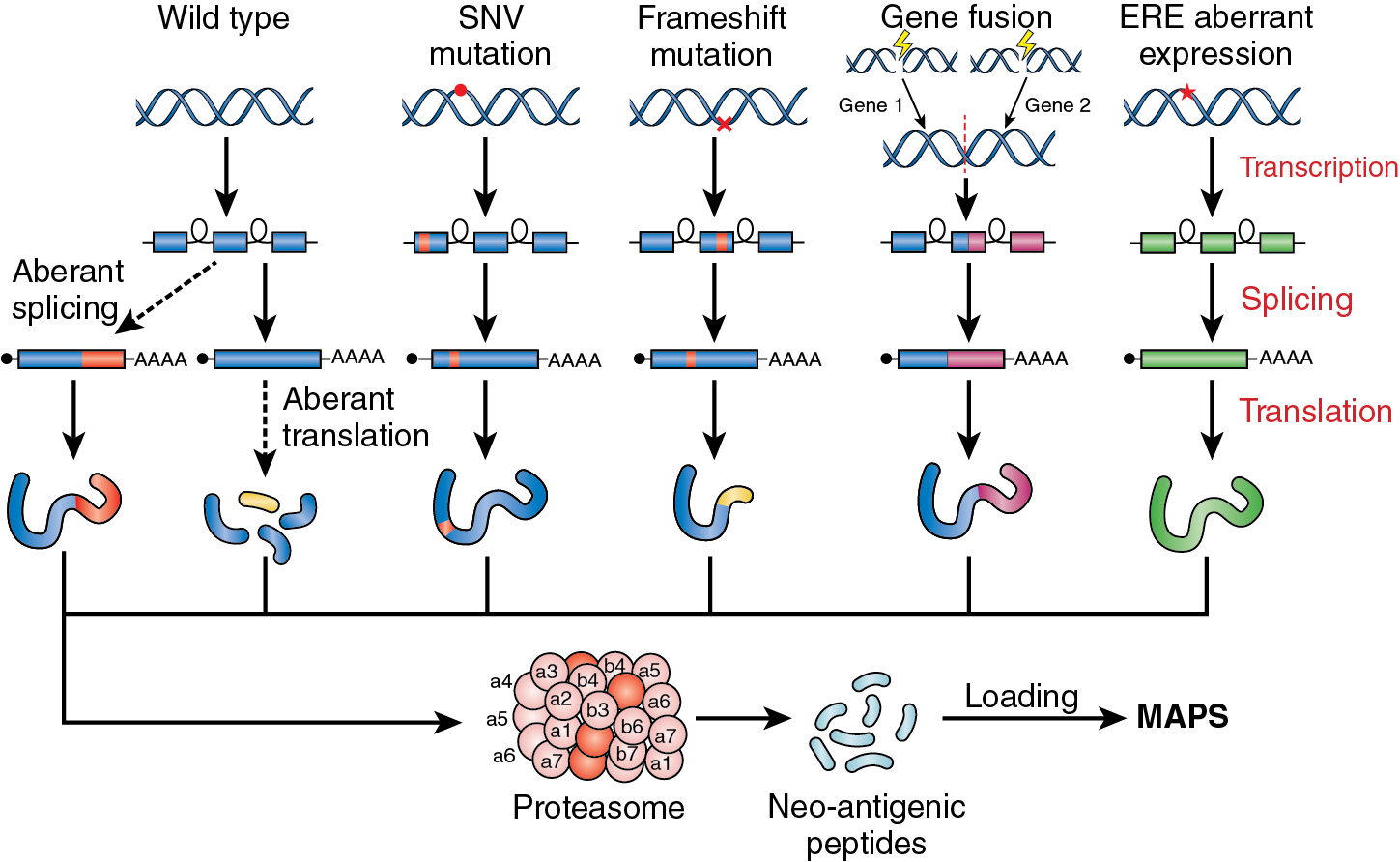
As discussed previously, the transcriptomes of cancer cells are highly vulnerable to dysregulation, which can also occur in the pre-mRNA stage on the path to gene transcription and final protein production. Dysregulation within the tumor cell at the pre-mRNA stage occurs via cis -acting mutations targeting the actual splicing of the exons. The highly regulated process of mRNA splicing in the pre-mRNA stage is a function of specific sequence of elements present on the pre-mRNA template, as well as many RNA processing proteins. This is often referred to as the supraspliceosome , and genetic alterations of the many factors involved in mRNA splicing may also provide a rich and expanding source of cancer-unique immunogenic proteins/peptides. Research reported by Seiler and colleagues demonstrated that 119 splice factor genes had significantly elevated rates of nonsynonymous mutations in a group of 33 different tumor types examined. There is increasing validation for aberrant splice factor genes as a rich source of highly antigenic epitopes capable, in animal models, of eliciting strong T cell–mediated antitumor responses. These observations are generating an interest in the induction of “forced” aberrant genetic alterations of highly selected splice factor genes as a novel therapeutic approach for augmenting immune checkpoint inhibition. The search for and characterization of these novel cancer-specific genetically altered splice factor genes has been greatly enhanced by new technologies such as high-throughput screening, Whole Genome Sequencing (WGS), RNAseq of both bulk tumor and single cells, and computational tools applicable for mining multiomic, publicly available cancer databases.
RNA dysregulation
Genetic instability and high mutation rates in progressing cancers provide an ideal opportunity for RNA screening dysregulation as a source of neoantigens capable of being targetable TSAs. These have been broadly subdivided into five categories as follows:
- (1)
Altered RNA expression in which overexpression is most common. In case of overexpression, the mRNA alteration in the tumor cells consistently causes the overproduction of a normal protein in the cancer cell compared with the noncancer cell. Examples include CD19 and GPC2.
- (2)
Alternative splicing is the process normal cells commonly use for creating diversity in the protein family of a single gene. In the genetically unstable cancer cell, mechanisms regulating alternative splicing are significantly enhanced and more open to mutation, including actual mutation events at splice junction sites. The resultant protein isoforms often function to enhance tumor cell proliferation and the processes of invasion through metastasis. These protein isoforms may also alter the tumor’s response to therapy. Of course, some of the alternatively spliced exons and altered splice junctions can also be a source of highly cancer-specific antigenic peptides. , , This source of peptides is felt to be seen by the host as more antigenically foreign than TSAs arising from SNVs. Non-sense-mediated mRNA decay is a normal cellular process that degrades mRNA transcripts that contain premature stop codons. , However, mutations in the non-sense-mediated mRNA decay pathway may alter this process, resulting in neoantigens being formed. ,
- (3)
Noncanonical splicing can occur in tumor cells producing unusual chimeric mRNA transcripts and, therefore, potentially novel tumor-specific proteins that are foreign to the host. These fusion transcripts may occur between segments of distant genes or through splicing of segments of adjacent genes. It should be noted that chimeric RNAs have also been found in embryonic stem cells and other noncancer tissues. Though the potential exists for the products of chimeric RNAs to be biomarkers of the cancer and potential antigenic targets, this is very preliminary work and will require considerable future studies for validation. Another class of RNAs also found in normal cells are RNAs that have formed a covalently linked closed loop. These closed-loop RNAs have been shown to also occur at the time of pre-mRNA splicing. Although circular RNAs lack a 5′ cap, they can use other methods for actual translation. , This certainly raises the possibility that such cancer-specific circular RNAs may be sufficiently genetically altered to be a source of antigenic peptides. To support such research, investigators have developed a database for circular RNAs found in cancer (MiOncoCirc). ,
- (4)
RNA editing has been noted earlier as a posttranscriptional editing process in normal cells and is another important contributor to protein diversity. There exist many opportunities in the genomically unstable cancer cell for genetic alterations to occur at the level of the enzymes operative in the editing process. RNA editing–derived peptides have been found to generate antigenic peptides that could combine with MHC class I molecules and be recognized by tumor-infiltrating T cells. Nevertheless, though an intriguing possibility, much future work is required to show that such antigenic peptides are suitable for development as therapeutic targets.
- (5)
A discussion of expressed transposable elements (TEs) seems appropriate for this section on aberrant transcription. TEs are DNA sequences that possess the ability to move from one location in the genome to another. TEs represent close to half the human genome and were first identified by Barbara McClintock of Cold Spring Harbor over 50 years ago. TEs can be divided into two major classes (although there are many subgroups in each class) based on the method used for transposition. Class I TEs require reverse transcription of an RNA intermediate into a cDNA copy in order to transpose. Class II TEs are complete in that they encode the protein transposase, allowing them to move and never use RNA intermediates. Dysregulation of TEs secondary to loss of DNA methylation is common in malignancy and permits reexpression of previously silenced TEs. Thus, apparently expressed TEs in cancer may result in proteins and their derived peptides that are antigenic to the host’s immune system. , , ,
Neoantigens resulting from coding sequence rearrangements (gene fusion TSAs)
These potential TSAs are relatively uncommon in cancer and result from the chromosomal rearrangement and juxtaposition of two independent DNA coding regions. These gene fusions may occur in nontumor cells, but many have been identified in a variety of cancers; for example, leukemias, sarcomas, breast cancer, and colon cancer. The most famous, of course, is the translocation that resulted in the formation of the BCR-ABL gene on chromosome 22 known as the Philadelphia chromosome in chronic myelogenous leukemia (CML) and other leukemias. Gene fusions occur by (1) chromosomal translocation, (2) transcriptional read-through of adjacent genes, or (3) trans – and cis -splicing of pre-mRNAs. The resultant fusion transcript can be translated into a novel fusion protein. Often these fusion or chimeric proteins are used clinically as biomarkers in diagnosis, response to therapy, and prognosis. ,
Recent evidence seems to highlight the importance of fusion peptides as effective tumor regression antigens. Investigators at Memorial Sloan Kettering Cancer Center identified a patient with metastatic head and neck cancer that had a complete response to immune checkpoint inhibitor therapy despite a tumor with a relatively low mutational load and minimal tumor-infiltrating T cells. Using whole-genome sequencing and RNA sequencing, they identified a novel gene fusion and showed that the resultant peptide neoantigen could elicit a host T-cell response. Using this approach, they identified a series of gene fusion–derived neoantigens that generated CTL responses.
Other fusion-driven cancers include EWS-FLI1 in Ewing sarcoma, EML4-ALK in lung adenocarcinoma, TMPRSS2-ERG in prostate cancer, ETV6-RUNX1 , FGFR3-TACC3 , TEL-AML1 , and others, including driver fusions, have been reported. Fusion proteins that function as oncogenic drivers appear to be highly clonal and thus it would not be advantageous to the tumor to reduce their expression. Clonality, however, they may occur only during the early initiation stages of the tumor process, becoming more heterogeneous during later stages, especially during metastatic development. In thinking about the future of research to identify oncogenic fusions, it would appear that fusions with breakpoints shared across tumor types and across patients would be highly relevant to developing vaccine and chimeric antigen receptor (CAR) T-cell therapies. There is evidence that these oncogenic fusions may be more relevant for tumors that have a relatively low tumor somatic mutational burden and a minimal immunocyte infiltration.
Endogenous nonmutated antigens
Endogenous retroviruses (ERVs) are a significant proportion of the human genome. The reference genome is reported to contain some 450,000 ERV-derived sequences. ERVs resulted from the integration into our genome of transposable elements more than 30 million years ago. As a result of this long evolutionary process, the vast majority of endogenous retroelements (EREs) in our genome are now truncated or mutated and can no longer transpose. It should be noted that ERVs are a major source of millions of gene regulatory elements as seen in pluripotent cells of early embryogenesis.
During the progression of cancer and the accompanying genomic instability, previous studies have found that dysregulation of certain EREs can lead to transcription and translation of aberrantly expressed EREs (aeEREs) generating viral-like neoantigenic peptides. It is possible that such nonmutated neoantigens could be shared TSAs across tumor types and hosts. During the early stages of cancer initiation and progression, induction of the viral mimicry response may occur, causing peptides translated from repeat-derived RNAs to be generated. These have been found to comprise the majority of the ERE-derived neoantigens processed and presented for immune recognition.
Neoantigens resulting from mutations in tumor HLA
Mutations that actually occur in the genes coding for the tumor MHC molecule, though apparently relatively uncommon, have become more readily identified. It is not unreasonable to accept that these somatically mutated MHC molecules could, in themselves, be a recognizable TSA. This occurrence was first described by Brandel et al. in 1996. A similar finding was also reported for melanoma, showing evidence that tumor-specific T cells could recognize antigens (polypeptide fragments of potential tumor antigens) presented in association with somatically mutated tumor HLA-A11.
It is possible that this form of TSA neoantigen is more common than initially thought because a number of new software tools have emerged that can identify HLA somatic mutations with a high degree of sensitivity (sensitivity 94.1%, specificity 53.3%). Using software tools such as NetMHCpan, it now appears possible to predict the sets of tumor antigenic peptides that would actually bind to the mutated MHC class I molecule and be recognized by the T-cell antigen receptor.
Posttranslational TSAs
Posttranslational modification of proteins within the cytoplasm is a biochemical process that adds specific biochemical moieties to specific protein residues. The most common biochemical posttranslational modification is phosphorylation. Others include sumoylation, acetylation, and ubiquitination. Studies have demonstrated that these modifications of proteins contribute to the malignant phenotype, including alterations in cell signaling, apoptosis, and transcriptional regulation. Though it is logical to assume that such posttranslation modified proteins can be recognized as foreign antigens and result in peptides suitable for binding to the cleft of MHC class I molecules, it has been challenging to confidently confirm such and to establish these as viable therapeutic antigen targets ( Box 5.1 ).