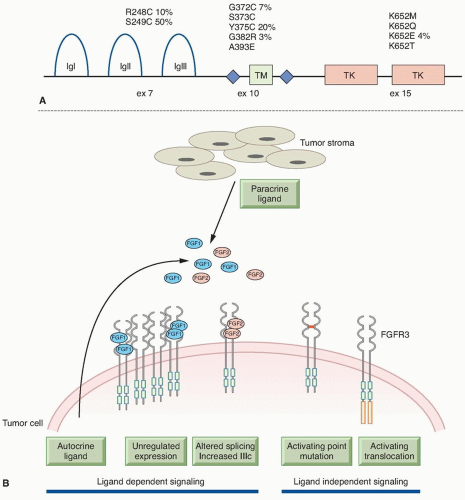
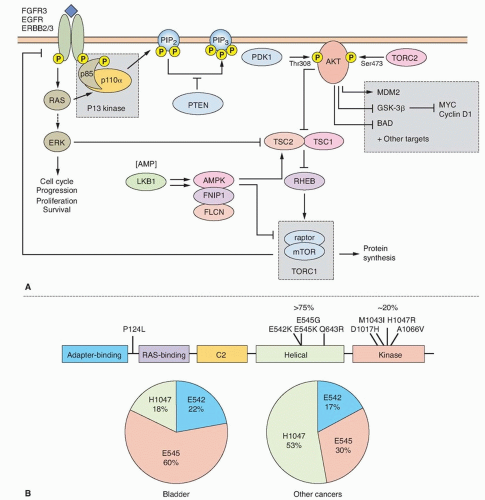
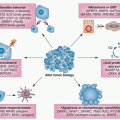
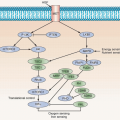
The mutation spectrum (Fig. 38.2B) differs significantly from that found in other cancers. Mutations E542K and E545K in the helical domain are most common (22% and 60%, respectively) and the kinase domain mutation H1047R, which is the most common mutation in other cancers, is less frequent. The selective pressure for helical domain mutation in UC is not fully understood. E542K and E545K forms require interaction with RAS-GTP but not binding to p85, the regulatory subunit of PI3K, whereas H1047R depends on p85 binding and is active in the absence of RAS binding.29 This suggests potential cooperation between helical domain PIK3CA mutant proteins and events in UC that activate RAS. Compatible with this, PIK3CA and FGFR3 mutations are commonly found together.25,26,27,28 Mutant PIK3CA confers a proliferative advantage at confluence and stimulates intraepithelial movement in NHUC, with higher activity of helical domain than kinase domain mutants in this cell context.30 Several other mechanisms of PI3K pathway activation have been identified in UC, though none of these are common in noninvasive tumors (Table 38.1).
cohesion between sister chromatids following DNA replication to ensure correct chromosomal segregation. Mutations in STAG2 in glioblastoma have been reported to cause aneuploidy35 but this relationship is not apparent in UC, where most mutations have been found in low grade/stage, genomically stable tumors, and there is no association of mutation with chromosomal copy number changes.31,34
TABLE 38.1 Genetic Changes Identified in Stage Ta Bladder Tumors | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
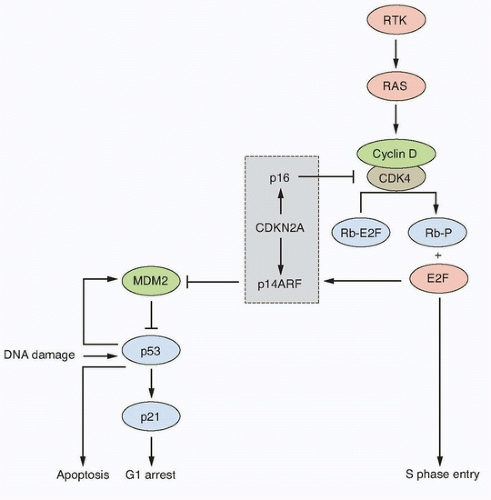
Inactivation of this locus in UC is commonly by homozygous deletion (HD). LOH of 9p, HD of CDKN2A, and loss of expression of p16 in NMI UC is predictive of reduced recurrence-free interval.50,51,52 Mouse knockout and in vitro experiments indicate that p16 and/or p14ARF may be haploinsufficient.53,54 In human UC, this may affect the biology of approximately 45% of cases that have LOH or reduced copy number of 9p21.
UC cells with epithelial phenotype have enhanced sensitivity to FGFR inhibition compared to those with mesenchymal phenotype.80 A recent RNAi screen in UC cell lines indicated that upregulated EGFR signaling provides a mechanism of escape from FGFR inhibition and can mediate de novo resistance. A reciprocal relationship was found when EGFR was inhibited and both in vitro and in a preclinical in vivo model, combined inhibition showed improved anti-tumor activity.81 Thus, assessment of both FGFR3 and EGFR status may be required to predict response and combined EGFR and FGFR3 inhibition may be essential. Several clinical trials of FGFR inhibitors are now planned or in progress in advanced UC.
TABLE 38.2 Genetic Changes Identified in Invasive (Stage ≥T2) Bladder Tumors | |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
prognostic significance of FGFR1 upregulation, and FGFR1 has not yet been examined independently of FGFR3 as a potential therapeutic target.
Stay updated, free articles. Join our Telegram channel

Full access? Get Clinical Tree